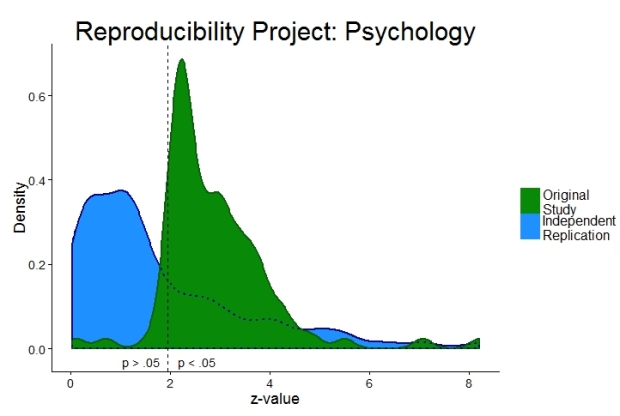
Psychological science is surprisingly difficult to replicate (Open Science Collaboration, 2015). Researchers are desperate to find out why. A new study in the prestigious journal PNAS (Van Bavel et al., 2016) claims that unknown contextual factors of psychological phenomena (“hidden moderators”) are to blame. The more an effect is sensitive to unknown contextual factors, the less likely a successful replication is. In this blog post I will lay out why I am not convinced by this argument.
Before I start I should say that I really appreciate that the authors of this paper make their point with reference to data and analyses thereof. I believe that this is a big improvement on the state of the replicability debate of a few years back when it was dominated by less substantiated opinions. Moreover, they share their key data and some analysis code, following good scientific practice. Still, I am not convinced by their argument. Here’s why:
1) No full engagement with the opposite side of the argument
Van Bavel et al.’s (2016) suggested influence of replication contexts on replication success cannot explain the following patterns in the data set they used (Open Science Collaboration, 2015):
a) replication effect sizes are mostly lower than original effect sizes. Effects might well “vary by [replication] context” (p. 2) but why the consistent reduction in effect size when replicating an effect?
b) internal conceptual replications are not related to independent replication success (Kunert, 2016). This goes directly against Van Bavel et al.’s (2016) suggestion that “conceptual replications can even improve the probability of successful replications” (p. 5).
c) why are most original effects just barely statistically significant (see previous blog post)?
I believe that all three patterns point to some combination of questionable research practices affecting the original studies. Nothing in Van Bavel et al.’s (2016) article manages to convince me otherwise.
2) The central result completely depends on how you define ‘replication success’
The central claim of the article is based on the correlation between one measure of replication success (subjective judgment by replication team of whether replication was successful) and one measure of the contextual sensitivity of a replicated effect. While the strength of the association (r = -.23) is statistically significant (p = .024), it doesn’t actually provide convincing evidence for either the null or the alternative hypothesis according to a standard Bayesian JZS correlation test (BF01 = 1). [For all analyses: R-code below.]
Moreover, another measure of replication success (reduction of effect size between original and replication study) is so weakly correlated with the contextual sensitivity variable (r = -.01) as to provide strong evidence for a lack of association between contextual sensitivity and replication success (BF01 = 12, notice that even the direction of the correlation is in the wrong direction according to Van Bavel et al.’s (2016) account).
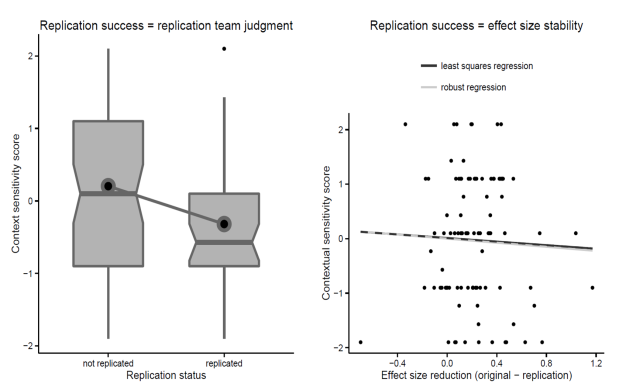
[Update: The corresponding values for the other measures of replication success are: replication p < .05 (r = -0.18; p = .0721; BF01 = 2.5), original effect size in 95%CI of replication effect size (r = -.3, p = .0032, BF10 = 6). I could not locate the data column for whether the meta-analytic effect size is different from zero.]
3) The contextual sensitivity variable could be confounded
How do we know which original effects were plagued by hidden moderators (i.e. by unknown context sensitivity) if, well, these moderators are hidden? Three of the authors of the article simply rated all replicated studies for contextual sensitivity without knowing each study’s replication status (but after the replication success of each study was known in general). The authors provide evidence for the ratings to be reliable but no one knows whether they are valid.
For example, the raters tried not to be influenced by ‘whether the specific replication attempt in question would succeed’ (p. 2). Still, all raters knew they would benefit (in the form of a prestigious publication) from a significant association between their ratings and replication success. How do we know that the ratings do not simply reflect some sort of implicit replicability doubt? From another PNAS study (Dreber et al., 2015) we know that scientists can predict replication success before a replication study is run.
Revealing hidden moderators
My problem with the contextual sensitivity account claiming that unknown moderators are to blame for replication failures is not so much that it is an unlikely explanation. I agree with Van Bavel et al. (2016) that some psychological phenomena are more sensitive to replication contexts than others. I would equally welcome it if scientific authors were more cautious in generalising their results.
My problem is that this account is so general as to be nearly unfalsifiable, and an unfalsifiable account is scientifically useless. Somehow unknown moderators always get invoked once a replication attempt has failed. All sorts of wild claims could be retrospectively claimed to be true within the context of the original finding.
In short: a convincing claim that contextual factors are to blame for replication failures needs to reveal the crucial replication contexts and then show that they indeed influence replication success. The proof of the unknown pudding is in the eating.
— — —
Dreber, A., Pfeiffer, T., Almenberg, J., Isaksson, S., Wilson, B., Chen, Y., Nosek, B., & Johannesson, M. (2015). Using prediction markets to estimate the reproducibility of scientific research Proceedings of the National Academy of Sciences, 112 (50), 15343-15347 DOI: 10.1073/pnas.1516179112
Kunert, R. (2016). Internal conceptual replications do not increase independent replication success Psychonomic Bulletin & Review DOI: 10.3758/s13423-016-1030-9
Open Science Collaboration (2015). Estimating the reproducibility of psychological science Science, 349 (6251) DOI: 10.1126/science.aac4716
Van Bavel, J.J., Mende-Siedlecki, P., Brady, W.J., & Reinero, D.A. (2016). Contextual sensitivity in scientific reproducibility PNAS
— — —
######################################################################################################## # Script for article "A critical comment on "Contextual sensitivity in scientific reproducibility"" # # Submitted to Brain's Idea # # Responsible for this file: R. Kunert (rikunert@gmail.com) # ######################################################################################################## # source functions if(!require(devtools)){install.packages('devtools')} #RPP functions library(devtools) source_url('https://raw.githubusercontent.com/FredHasselman/toolboxR/master/C-3PR.R') in.IT(c('ggplot2','RColorBrewer','lattice','gridExtra','plyr','dplyr','httr','extrafont')) if(!require(BayesMed)){install.packages('BayesMed')} #Bayesian analysis of correlation library(BayesMed) if(!require(Hmisc)){install.packages('Hmisc')} #correlations library(Hmisc) if(!require(reshape2)){install.packages('reshape2')}#melt function library(reshape2) if(!require(grid)){install.packages('grid')} #arranging figures library(grid) #get raw data from OSF website info <- GET('https://osf.io/pra2u/?action=download', write_disk('rpp_Bevel_data.csv', overwrite = TRUE)) #downloads data file from the OSF RPPdata <- read.csv("rpp_Bevel_data.csv")[1:100, ] colnames(RPPdata)[1] <- "ID" # Change first column name #------------------------------------------------------------------------------------------------------------ #2) The central result completely depends on how you define 'replication success'---------------------------- #replication with subjective judgment of whether it replicated rcorr(RPPdata$ContextVariable_C, RPPdata$Replicate_Binary, type = 'spearman') #As far as I know there is currently no Bayesian Spearman rank correlation analysis. Therefore, use standard correlation analysis with raw and ranked data and hope that the result is similar. #parametric Bayes factor test bf = jzs_cor(RPPdata$ContextVariable_C, RPPdata$Replicate_Binary)#parametric Bayes factor test plot(bf$alpha_samples) 1/bf$BayesFactor#BF01 provides support for null hypothesis over alternative #parametric Bayes factor test with ranked data bf = jzs_cor(rank(RPPdata$ContextVariable_C), rank(RPPdata$Replicate_Binary))#parametric Bayes factor test plot(bf$alpha_samples) 1/bf$BayesFactor#BF01 provides support for null hypothesis over alternative #replication with effect size reduction rcorr(RPPdata$ContextVariable_C[!is.na(RPPdata$FXSize_Diff)], RPPdata$FXSize_Diff[!is.na(RPPdata$FXSize_Diff)], type = 'spearman') #parametric Bayes factor test bf = jzs_cor(RPPdata$ContextVariable_C[!is.na(RPPdata$FXSize_Diff)], RPPdata$FXSize_Diff[!is.na(RPPdata$FXSize_Diff)]) plot(bf$alpha_samples) 1/bf$BayesFactor#BF01 provides support for null hypothesis over alternative #parametric Bayes factor test with ranked data bf = jzs_cor(rank(RPPdata$ContextVariable_C[!is.na(RPPdata$FXSize_Diff)]), rank(RPPdata$FXSize_Diff[!is.na(RPPdata$FXSize_Diff)])) plot(bf$alpha_samples) 1/bf$BayesFactor#BF01 provides support for null hypothesis over alternative #------------------------------------------------------------------------------------------------------------ #Figure 1---------------------------------------------------------------------------------------------------- #general look theme_set(theme_bw(12)+#remove gray background, set font-size theme(axis.line = element_line(colour = "black"), panel.grid.major = element_blank(), panel.grid.minor = element_blank(), panel.background = element_blank(), panel.border = element_blank(), legend.title = element_blank(), legend.key = element_blank(), legend.position = "top", legend.direction = 'vertical')) #Panel A: replication success measure = binary replication team judgment dat_box = melt(data.frame(dat = c(RPPdata$ContextVariable_C[RPPdata$Replicate_Binary == 1], RPPdata$ContextVariable_C[RPPdata$Replicate_Binary == 0]), replication_status = c(rep('replicated', sum(RPPdata$Replicate_Binary == 1)), rep('not replicated', sum(RPPdata$Replicate_Binary == 0)))), id = c('replication_status')) #draw basic box plot plot_box = ggplot(dat_box, aes(x=replication_status, y=value)) + geom_boxplot(size = 1.2,#line size alpha = 0.3,#transparency of fill colour width = 0.8,#box width notch = T, notchwidth = 0.8,#notch setting show_guide = F,#do not show legend fill='black', color='grey40') + labs(x = "Replication status", y = "Context sensitivity score")#axis titles #add mean values and rhythm effect lines to box plot #prepare data frame dat_sum = melt(data.frame(dat = c(mean(RPPdata$ContextVariable_C[RPPdata$Replicate_Binary == 1]), mean(RPPdata$ContextVariable_C[RPPdata$Replicate_Binary == 0])), replication_status = c('replicated', 'not replicated')), id = 'replication_status') #add mean values plot_box = plot_box + geom_line(data = dat_sum, mapping = aes(y = value, group = 1), size= c(1.5), color = 'grey40')+ geom_point(data = dat_sum, size=12, shape=20,#dot rim fill = 'grey40', color = 'grey40') + geom_point(data = dat_sum, size=6, shape=20,#dot fill fill = 'black', color = 'black') plot_box #Panel B: replication success measure = effect size reduction dat_corr = data.frame("x" = RPPdata$FXSize_Diff[!is.na(RPPdata$FXSize_Diff)], "y" = RPPdata$ContextVariable_C[!is.na(RPPdata$FXSize_Diff)])#plotted data plot_corr = ggplot(dat_corr, aes(x = x, y = y))+ geom_point(size = 2) +#add points stat_smooth(method = "lm", size = 1, se = FALSE, aes(colour = "least squares regression")) + stat_smooth(method = "rlm", size = 1, se = FALSE, aes(colour = "robust regression")) + labs(x = "Effect size reduction (original - replication)", y = "Contextual sensitivity score") +#axis labels scale_color_grey()+#colour scale for lines stat_smooth(method = "lm", size = 1, se = FALSE, aes(colour = "least squares regression"), lty = 2) plot_corr #arrange figure with both panels multi.PLOT(plot_box + ggtitle("Replication success = replication team judgment"), plot_corr + ggtitle("Replication success = effect size stability"), cols=2)